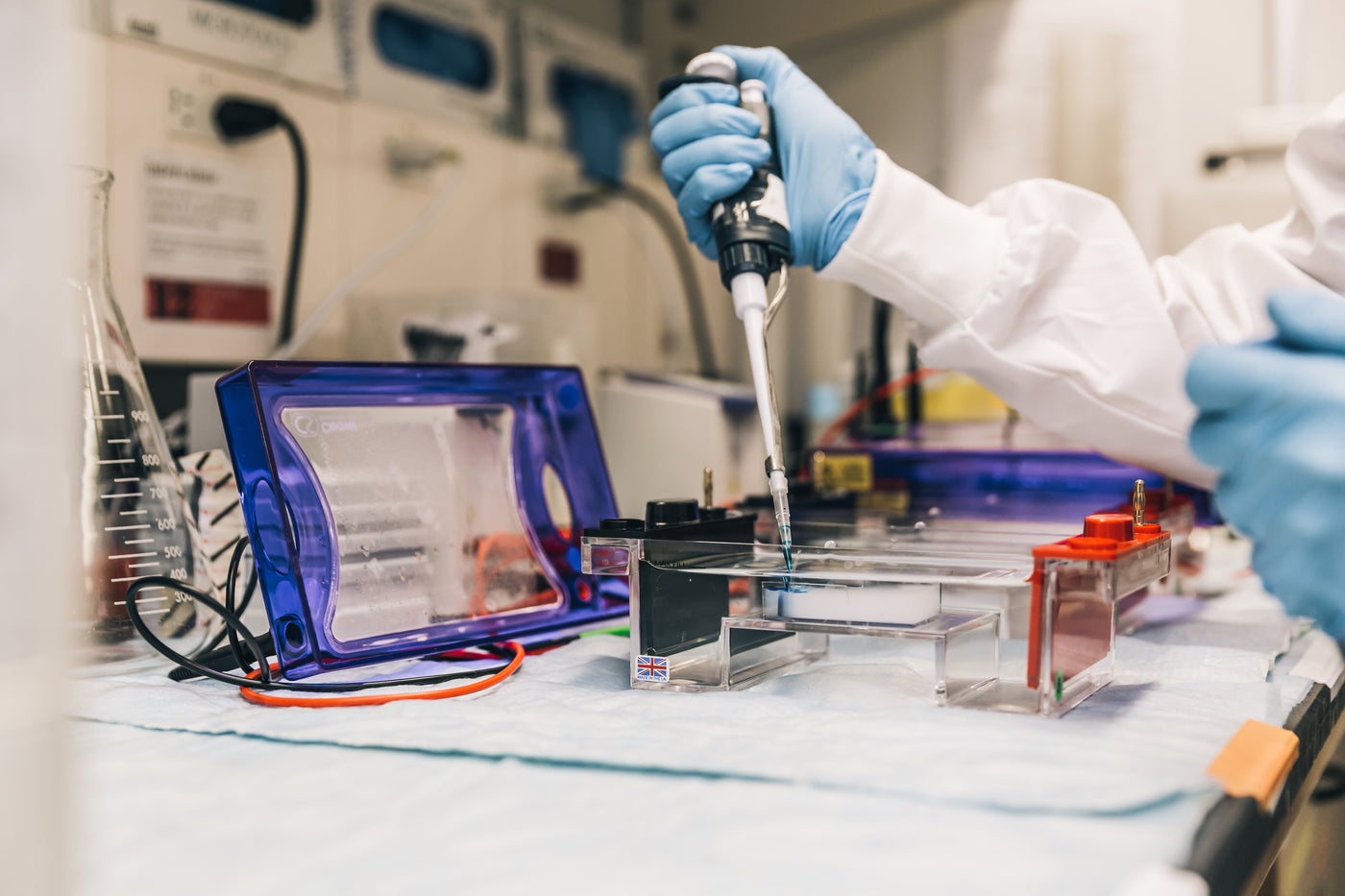
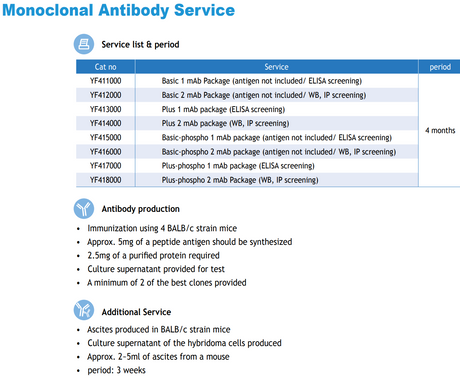
Your custom project will be ready in:
Order now custom recombinants
Featured collection
View allA headline to grab attention
Use this card with an image to highlight your collections or introduce a promotion.
Product features
Use this section to highlight specific product features to your customers.
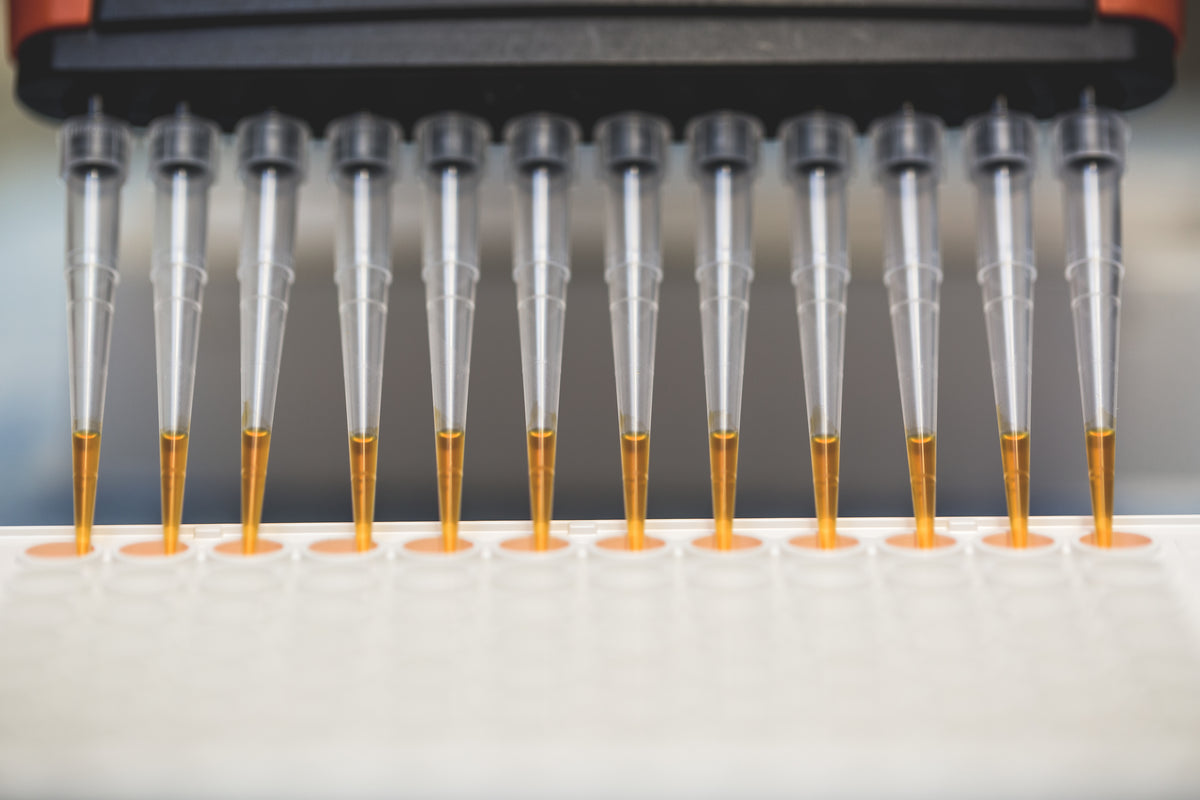

Media with text
Share information about your brand with your customers. Describe a product, make announcements, or welcome customers to your store.
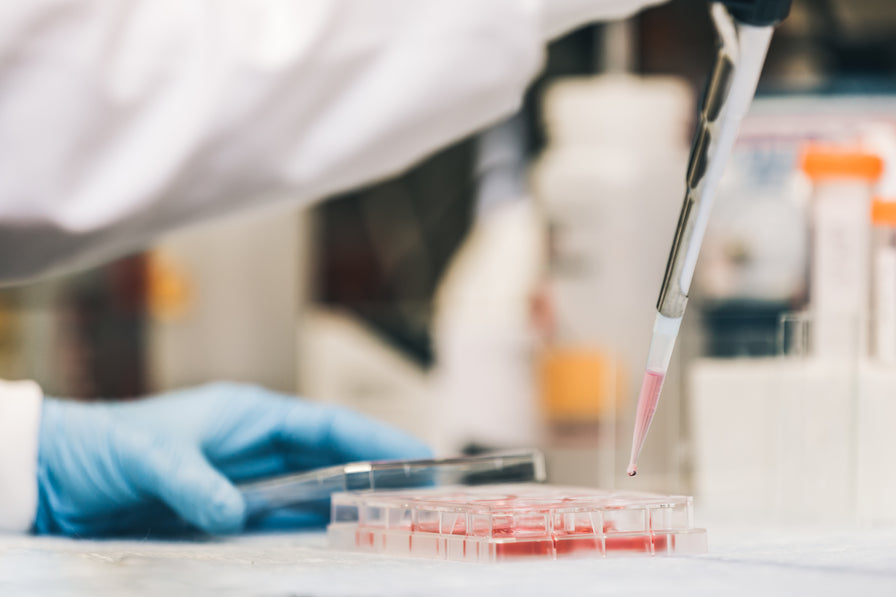
Shoppable image
Create a Shop the Look section to assist with upselling or helping customers with purchasing complementary products.
Link lists
Inventory in Europe and USA
(AXS-1114544) Lymphoprep - 250ml - 1 vial is backordered and will ship as soon as it is back in stock.
Product comparison grid
Add content here to explain a bit about the range of products on offer and which ones may be most suitable for your customers.
Multi-column
View AllVideo
Cellular ingestion